srukf
Run one update of a square-root unscented (sigma point) Kalman filter. The advantage of a "square-root" UKF over a traditional UKF is that the covariance can be maintained in a more numerically stable way.
[x_k, S_k] = srukf( ...
t_km1, t_k, x_km1, S_km1, u_km1, z_k, ...
f, h, sqQ_km1, sqR_k, ...
alpha, beta, kappa, ...
(etc.))Inputs
t_km1 | Time at sample k-1 |
|---|---|
t_k | Time at sample k |
x_km1 | State estimate at sample k-1 |
S_km1 | Cholesky factor (lower diagonal) of the estimate covariance
at sample k-1, such that the covariance is |
u_km1 | Input vector at sample k-1 |
z_k | Measurement at sample k |
f | Propagation function with interface: where |
h | Measurement function with interface: where |
sqQ | Matrix square root of the pocess noise covariance at k-1 |
sqR | Matrix square root of the measurement noise covariance at k |
alpha | Optional tuning parameter, often 0.001 |
beta | Optional tuning parameter, with 2 being optimal for Gaussian estimation error |
kappa | Optional tuning parameter, often 3 - |
(etc.) | Additional arguments to be passed to f and h after their normal arguments |
Outputs
x | Upated estimate at sample k |
|---|---|
S | Updated Cholesky factor of the estimate covariance at k |
Example
We can quickly create a simulation for discrete, dynamic system, generate noisy measurements of the system over time, and pass these to a square-root unscented Kalman filter.
First, define the discrete system.
rng(1);
dt = 0.1; % Time step
F_km1 = expm([0 1; -1 0]*dt); % State transition matrix
H_k = [1 0]; % Observation matrix
G_km1 = [0.5*dt^2; dt]; % Process-noise-to-state map
Q_km1 = G_km1 * 0.5^2 * G_km1.'; % Process noise variance
R_k = 0.1; % Meas. noise covarianceThe srukf algorithm uses the sqrts of the process and measurement noise
as well. These can be a cholesky factor or the form returned by
sqrtpsdm; it doesn't matter.
sqQ = sqrtpsdm(Q_km1);
sqR = sqrtpsdm(R_k);Make propagation and observation functions. These are just linear for
this example, but srukf is meant for nonlinear functions.
f = @(t_km1, t_k, x_km1, u_km1, q_km1) F_km1 * x_km1 + q_km1;
h = @(t_k, x_k, u_km1, r_k) H_k * x_k + r_k;Now, we'll define the simulation's time step and initial conditions. Note that we define the initial estimate and set the truth as a small error from the estimate (using the covariance).
n = 100; % Number of samples to simulate
x_hat_0 = [1; 0]; % Initial estimate
P = diag([0.5 1].^2); % Initial estimate covariance
S = sqrtpsdm(P, 'L'); % Initial sqrt of the covariance
x_0 = x_hat_0 + mnddraw(P, 1); % Initial true stateNow we'll just write a loop for the discrete simulation.
% Storage for time histories
x = [x_0, zeros(2, n-1)]; % True state
x_hat = [x_hat_0, zeros(2, n-1)]; % Estimate
z = [H_k * x_0 + mnddraw(R_k, 1), zeros(1, n-1)]; % Measurement
% Simulate each sample over time.
for k = 2:n
% Propagate the true state.
x(:, k) = F_km1 * x(:, k-1) + mnddraw(Q_km1, 1);
% Create the real measurement at sample k.
z(:, k) = H_k * x(:, k) + mnddraw(R_k, 1);
% Run the Kalman correction.
[x_hat(:,k), S] = srukf((k-1)*dt, k*dt, x_hat(:,k-1), S, [], z(:,k),...
f, h, sqQ, sqR);
endPlot the results.
figure(1);
clf();
t = 0:dt:(n-1)*dt;
plot(t, x, ...
t, z, '.', ...
t, x_hat, '--');
legend('True x1', 'True x2', 'Meas.', 'Est. x1', 'Est. x2');
xlabel('Time');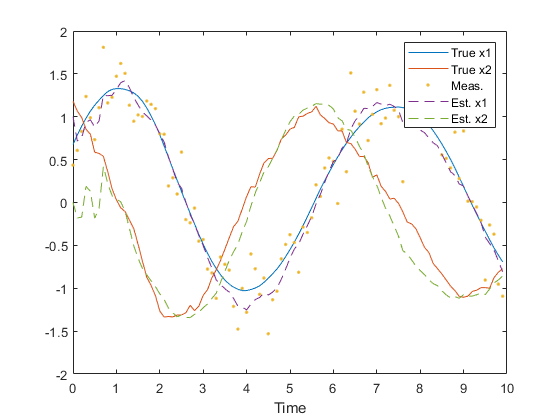
Note how similar this example is to the example of ukf.
Reference
Wan, Eric A. and Rudoph van der Merwe. "The Unscented Kalman Filter." Kalman Filtering and Neural Networks. Ed. Simon Haykin. New York: John Wiley & Sons, Inc., 2001. Print. Pages 273-275.
See Also
*kf v1.0.3 January 17th, 2025
©2025 An Uncommon Lab